1-6 of 6 results
-
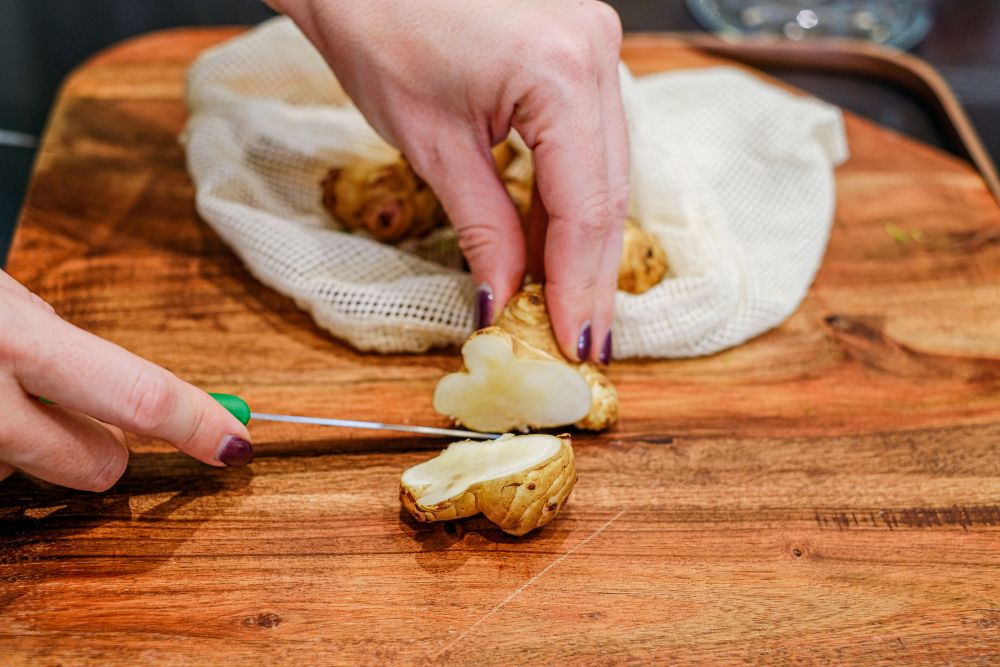
What happens when you eat fiber, and why you should eat more
In recent years much research has been done determining that the products of a healthy gut microbiota also have health benefits way beyond the gut. -
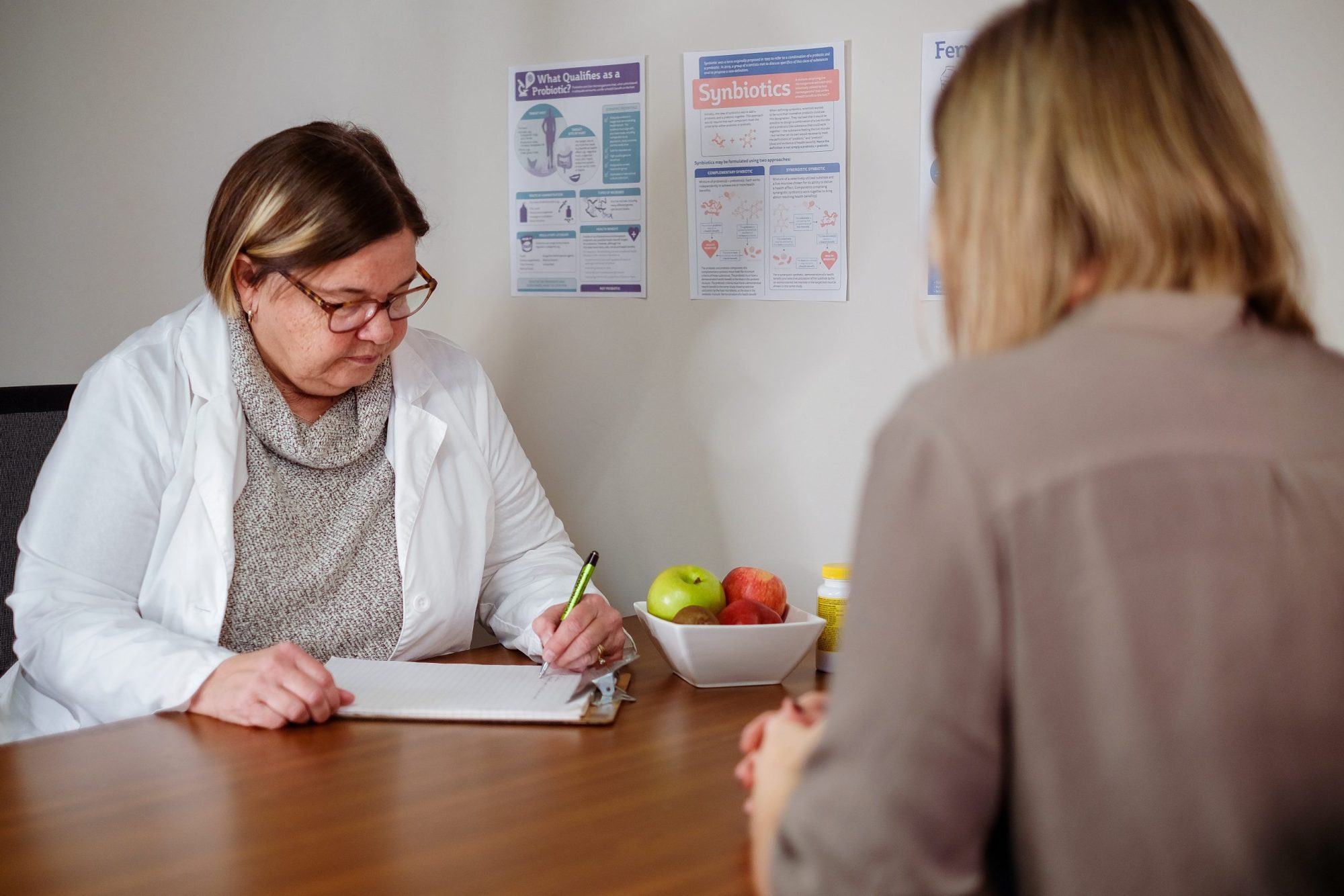

Planning a Biotics Study? New Publication Recommends Adding Diet as a Variable
Both dietary patterns and specific foods are known to affect the digestive tract environment, either directly or indirectly (for example, by shaping the gut microbiome), and are therefore of interest as potential factors in the response of an individual to a biotic intervention. -
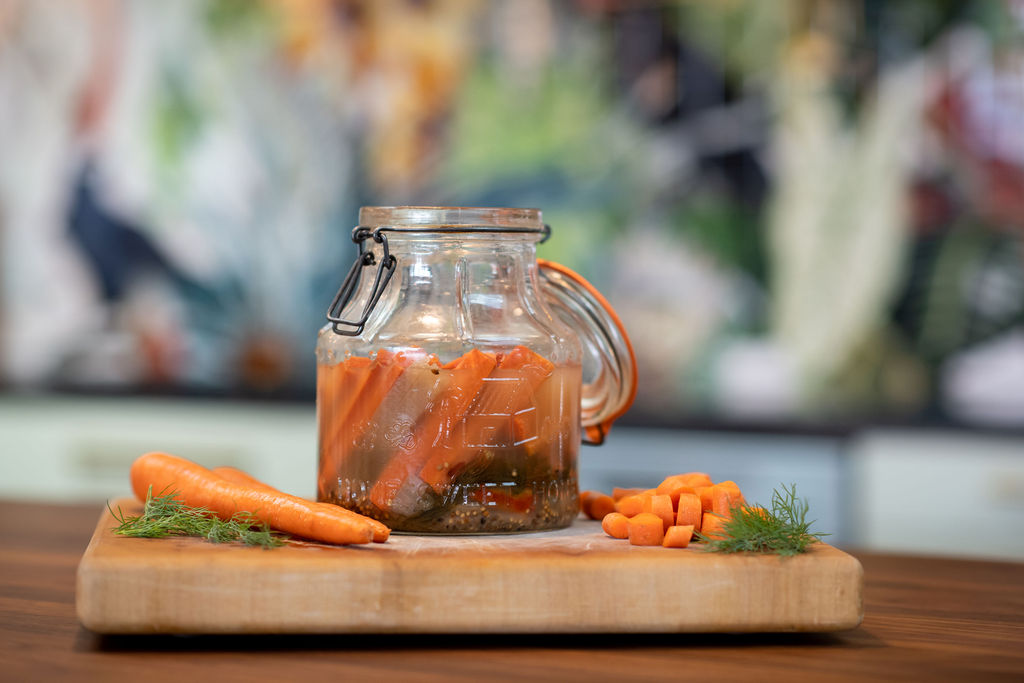

Food of the future: Fermented and sustainable
By Dr. Mary Ellen Sanders, ISAPP Executive Science Officer An exciting research initiative at the crossroads of fermented foods and… -

Bifidobacteria in the infant gut use human milk oligosaccharides: how does this lead to health benefits?
By Martin Frederik Laursen, Technical University of Denmark, 2022 co-recipient of Glenn Gibson Early Career Research Prize Breast milk is… -
The Human Mycobiome: An ISAPP mini-symposium
ISAPP announces an open registration mini-symposium on the human mycobiome. Although the contribution of the intestinal microbiome in human physiology… -
I have IBS – should I have my microbiome tested?
By Prof. Eamonn Quigley, MD. The Methodist Hospital and Weill Cornell School of Medicine, Houston I am a gastroenterologist and…
