1-11 of 11 results
-

New ISAPP Webinar: Fermented Foods and Health — Continuing Education Credit Available for Dietitians
Dietitians – along with many other nutritional professionals – often receive questions about consuming fermented foods for digestive health. But… -

ISAPP board members give a scientific overview of synbiotics in webinar
Many kinds of products are labeled as synbiotics – but how do they differ from each other? And do they… -
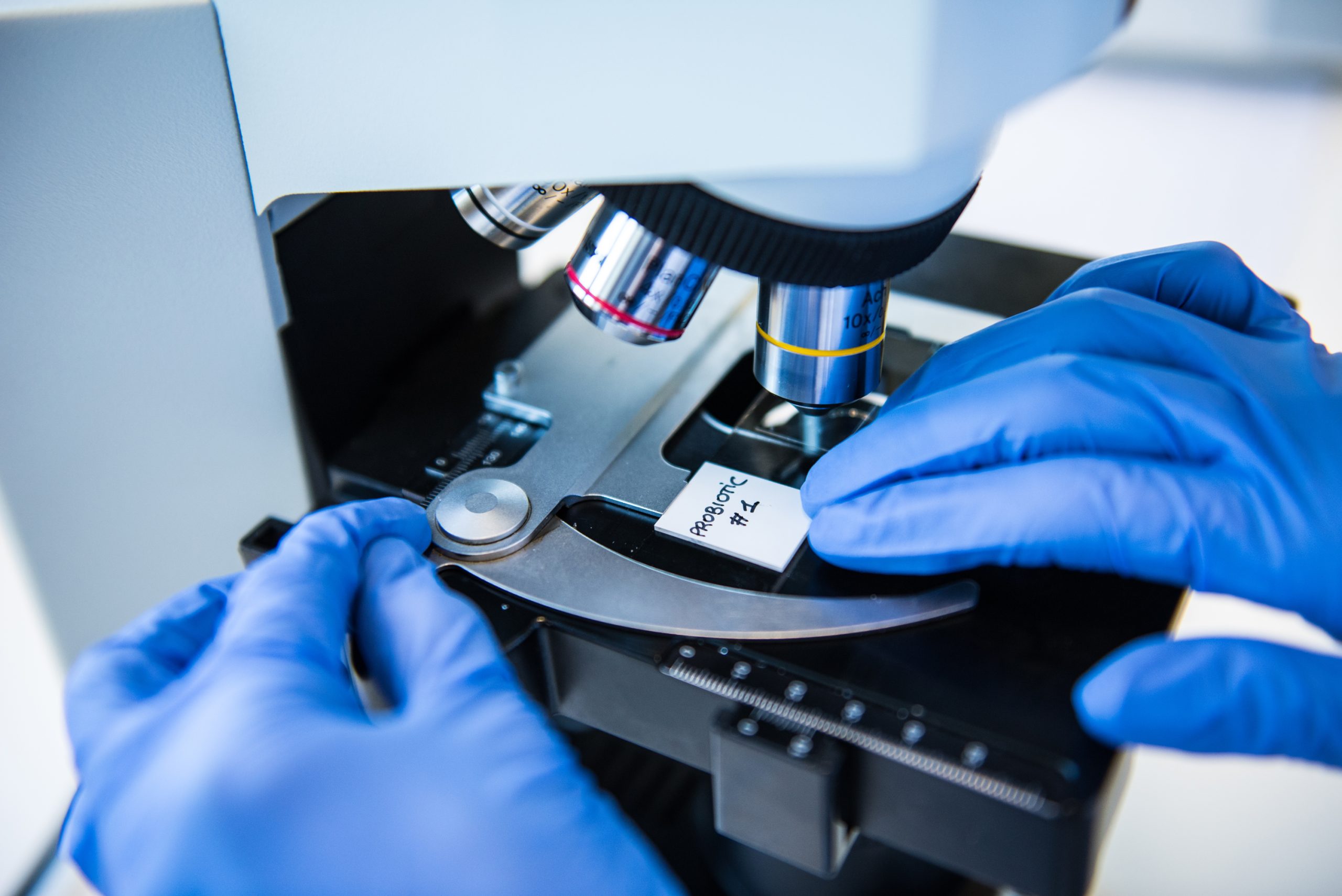

Hear from ISAPP board members in webinar covering probiotic and prebiotic mechanisms of action
This webinar is now complete — see the recorded version here. New probiotic and prebiotic trials are published all the… -
ISAPP partners with British Nutrition Foundation for fermented foods webinar
Did you miss the live webinar? Access the archived version here. Read the speaker Q&A here. From sourdough starter tips… -

Limitations of microbiome measurement: Prof. Gloor shares insights with ISAPP
February 20, 2019 The number of papers published on the human microbiome is growing exponentially – but not all of… -

ISAPP conducts webinar on definitions in microbiome space for ILSI-North America Gut Microbiome Committee
Dr. Mary Ellen Sanders presented a webinar July 23, 2018 – covering basic definitions of microbiota-mediated terminology – to the… -

ISAPP’s Outgoing President: Karen Scott
Dr. Karen Scott of the Rowett Institute of the University of Aberdeen has served as the ISAPP President for the last three… -
Medscape Webinar on Probiotics – Now Available!
“Navigating the world of probiotics: Helping patients make good choices” This 30-min CME activity, which took place on April 17th,… -
Medscape Webinar on Probiotics April 17
Prof. Dan Merenstein MD and Mary Ellen Sanders, PhD will present a 30 min webinar titled, “Navigating the World of… -
ISAPP to host live webinar: Microbial metabolism associated with health
Update April 16, 2018: Recording and slides from the webinar available here. The International Scientific Association for Probiotics and Prebiotics (ISAPP), in… -

Two Free Webinars on Probiotics!
Both webinars – eligible for continuing education credit – on probiotics involving ISAPP board members are scheduled. The first is…